Flexible Multi-level Models for Longitudinal Analysis of Childhood Obesity
Research Question
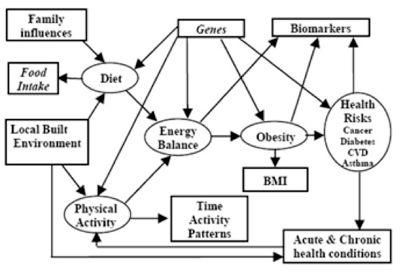
Guided by a conceptual pathway for obesity (given in the figure), the project has the following three main aims:
Aim 1: To develop flexible and functional-based multi-level quantile regression techniques for longitudinal modeling of childhood obesity via:
1a) assessment of effects of diet, physical activity, built-environment, demographic, psycho-social and developmental factors (at various levels of aggregation) on individual BMI trajectories directly, or relative to age- and gender- specific percentile reference curves,
1b) extension of the multi-level, longitudinal quantile regression framework to incorporate hierarchical modeling to investigate patterns of quantile-specific effects and to include prior covariates describing characteristics among risk factors.
Aim 2: To develop new methods for the analysis of genes and gene-environment (GxE) interactions using family data in the context of longitudinal growth curves:
2a) a longitudinal version of the quantitative transmission disequilibrium test (the L-QTDT) for the analysis of repeated BMI measurements in a parent-offspring trio sample; also allowing for inference in the quantile regression setup
2b) a method to find obesity-related genes involved in a GxE interaction (e.g. genes that modify the effect of dietary intake) in the context of a candidate-gene or genomewide association study.
Aim 3: To develop new latent-variable and associated structural equation modeling approaches
3a) to allow for mediational effects (e.g., diet) and unobserved latent variables (e.g., energy balance) in the flexible multi-level quantile regression models proposed in Aim 1; and
3b) to develop an integrated modeling framework for jointly assessing the effects of the built environment, biomarkers (e.g., leptin) potentially available on a subset cohort of children, and genes (e.g., FTO) on development of obesity during childhood.
Modeling Approach
Dr. Berhane has continued to work with a PhD student Roger Chang on new functional based quantile regression multi-level models (Aim 1a). They have successfully implemented a new model via a Monte Carlo Expectation-Maximization (MCEM) approach to allow for multiple random effects by extending previous relatively simpler mixed effects quantile regression approaches that only allow for a random intercept. Drs. Conti and Berhane are working with a PhD student (Ernest Shen) on estimating a complete quantile process through use of hierarchical modeling techniques (Aim 1b). A second-stage estimation of the conditional quantiles is conducted by specifying a second-stage linear model. This model provides a Wald test for model specification and allows for improved estimation of model parameters across the entire quantile process. Future work will: a) develop spatial statistical methods for estimation b) consider non-linear second-stage model specifications; and c) enrich the second-stage model by including other covariates that may influence on the first-stage parameter estimates. The work will develop frequentist approaches with more sophisticated estimation procedures for the second-stage. We extended hierarchical modeling by incorporating model selection showing that a well-defined phenotype-specific set of priors reduces the model search space and enhances biological relevance to final inference. This work has now been extended for analysis of rare genetic variants, as well (Quintanta et al. submitted).
Dr. Gauderman (with a PhD student Ray Su) worked (Aim 2) on a comprehensive review of pathway approaches by simulating data from CHS; with a goal to develop a novel pathway approach that could have improvement of power compared with current methods. In addition, the goal is to develop a standardized comparison ground among competitive and self-contained tests. We will apply this pathway approach to investigate BMI associations in a genetics framework under the Children Health's Study. Dr. Thomas published three reviews of methods for GxE interactions (Thomas, 2010ab; Thomas et al. (2009)) relating to the genetic (Aim 2) and integrated modeling (Aim 3) of the study. The Nature Reviews Genetics (Thomas, 2010b) in particular draws upon examples from the Children's Health Study for examples of gene-environment. Two more recent papers (Baurley et al., 2010; Chen and Thomas, 2010) have described novel methods for screening for higher-order interactions, which are scalable to genome-wide studies. Another contribution (Wilson et al., 2010) describes Bayesian methods for fitting complex models like those we are developing in Aim 3a.
Drs. Chou and Berhane are working with a PhD student, Ernest Shen, to develop techniques for estimating conditional quantiles when dealing with mediation analysis. This work extends existing methods on mediation models that have focused on estimation of conditional means for such path models. Work on this topic follows two related strains: the first follows up on the point that the total effect of a predictor X on response Y is equal to the sum of the direct effect and the indirect effect of X on Y as mediated through another variable, say M. We checked whether this identity holds across the whole response distribution via simulation, particularly at the extreme tails. Preliminary results indicate that the conditional mean mediation effects are quite different from those given by the tails. We have derived Wald tests for the conditional quantile mediation effects. We are examining whether previous conditional mean methods have the same finite-sample properties for a range of conditional quantiles, and results suggest that Wald tests near the tails of the distribution (e.g. 95th conditional quantile effects at which data sparsity is an issue) have substantially less power than those conducted for central quantiles. We will establish the conditions under which the identity can be proven analytically. Dr. Chou examined piecewise growth curve model drug use trajectories of 14 waves of longitudinal data across four developmental stages (presented in Society of Prevention Research 2010 annual conference), showing that initial statuses and growth trajectories were significantly correlated across developmental stages. Chou, et. al. (2010) applied growth curve models to randomized trials, finding that the initial status as a parameter of the growth curve model is more meaningful the starting point of the post-test instead of the pre-test measures.
Drs. Thomas and Berhane have continued their work on Aim 3b, with focus on investigating the properties of a basic integrated model where a biomarker of the intermediate disease process is incorporated, via latent variable representing airway inflammation. In such models, it is of interest to compare estimation of the quantity of interest, the net effect of E on Y, in a basic integrated model to a standard regression model relating E to Y. In the case where all variables are normally distributed, through simulation and likelihood theory it has been shown that the net effect of E on Y and the variance of the estimate of the net effect in the integrated model are the same as for the standard regression model (to be presented at the Joints Statistical Meetings in August, 2011). A series of up to four manuscripts are currently in progress. Dr. Pentz and colleagues have made substantial progress in their research productivity and data collection for the healthy Places study, on which we will rely for substantive inspirations and illustrative applications for our newly developed methods. Dr. Pentz has been working closely with Drs. Chou and Berhane, who are also investigators in the Healthy Places Study.
Significance
The methods being developed are important in understanding effects of environmental and genetic factors on childhood obesity, beyond what is possible with existing methods. The involvement of several investigators in the related epidemiologic studies will enable the implementation and application of new methods to data from the CHS and the Healthy Places study. We expect this to have far reaching policy implication in preventing childhood obesity. We also expect that the new techniques will have significant impact on our ability to analyze data from studies with designs similar to those of the CHS and the Healthy Places study.
Principal Investigator
Kiros Berhane, PhD
Professor
Division of Biostatistics, Department of Preventive Medicine, Keck School of Medicine
University of Southern California
Co-Investigators
David Conti, PhD
Associate Professor
USC
Chih-Ping Chou, PhD
Professor
USC
W. James Gauderman
Professor
USC
Frank Gilliland, MD PhD
Professor
USC
Michael Goran, PhD
Professor
USC
Maryann Pentz, PhD
Professor
USC
Fredrick Schumacher, PhD
Assistant Professor
USC
Duncan Thomas, PhD
Professor
USC